Beyond DNA: Why Multi-Omic and Spatial Data Are Essential for Precision Cardiovascular Medicine
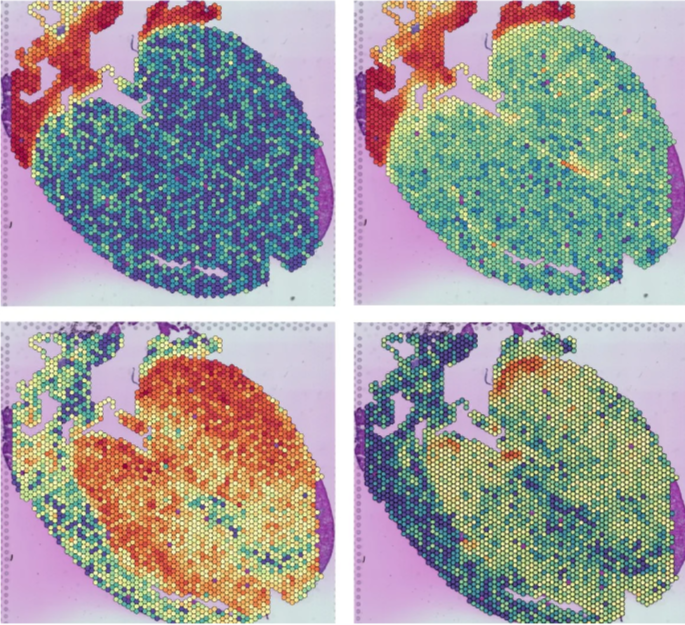
Differential gene expression patterns in cardiac tissue visualised using spatial transcriptomics.
Image source: Martin Heart Lab.
Author: Bahar Folad, Multi-omics Research Analyst, AXIA Medicine
Published by AXIA Medicine | February 2026
When Structural Precision Misguides Cardiovascular Care
Cardiovascular disease (CVD) remains the leading cause of death worldwide, accounting for roughly 30% of global mortality. Despite this burden, routine clinical care continues to rely primarily on anatomical imaging, haemodynamic measurements, and population-based risk scores. These approaches were highly effective in earlier eras of cardiovascular care, when disease models focused primarily on late-stage structural pathology and therapeutic options were relatively uniform. As disease understanding and therapeutic complexity have expanded, however, structural assessment alone provides an incomplete view of the biological processes driving progression and treatment response. Consequently, clinicians often face uncertainty in early risk resolution, treatment selection, and identification of patients most likely to benefit from targeted interventions.
Two patients with similar anatomical findings can follow markedly different clinical trajectories, reflecting molecular heterogeneity that conventional clinical measures cannot capture. This contributes to variable treatment responses and heterogeneity in clinical trials. Such heterogeneity dilutes measurable treatment effects, increases sample size requirements, and contributes to inconclusive trial outcomes that obscure subgroup benefit and delay therapeutic translation (Ramón y Cajal et al., 2020). Precision cardiovascular medicine therefore requires biological insight beyond anatomy and single molecular measurements. Multi-omic and multi-modal data enable understanding of cardiovascular disease at the level of cells, pathways, and dynamic physiological change.
Genomics as a Foundation but Not a Complete Solution
Genomics has advanced cardiovascular risk prediction and prevention, but primarily captures static inherited susceptibility rather than the dynamic molecular activity driving disease progression and treatment response (Wang, Maron, and Loscalzo, 2024). DNA-based markers cannot fully reflect tissue-specific gene expression, cellular context, or ongoing disease activity. Evidence from network medicine shows that cardiovascular risk variants operate within specific cellular environments such as endothelial cells, fibroblasts, cardiomyocytes, and immune populations. These cell-type–specific programmes shape inflammation, fibrosis, and vascular remodelling, processes that influence clinical presentation and progression but are not resolved through genomics alone. Genomics therefore provides essential background risk information but insufficient resolution for treatment selection or dynamic monitoring.
Similarly, bulk tissue and circulating biomarkers provide population-averaged signals that can mask biologically relevant changes within small or rare cell populations. This signal dilution may obscure patient-specific disease mechanisms, creating disconnects between trial outcomes and true therapeutic potential. The challenge is especially pronounced in heterogeneous conditions such as atherosclerosis, cardiomyopathies, and pulmonary vascular disease, where it limits precise risk stratification, therapeutic targeting, and dynamic monitoring (Ord et al., 2021; Jeong et al., 2025; Massad et al., 2025). In carotid artery stenosis, diagnosis and management remain dominated by anatomical imaging and haemodynamic measurements, capturing luminal narrowing but not the molecular pathways driving plaque instability or treatment response.
Integrated transcriptomic, proteomic, and metabolomic analyses reveal disease mechanisms and endotypes that anatomy alone cannot predict, enabling more precise patient stratification, intervention selection, and longitudinal monitoring (Liu et al., 2021; Henry et al., 2022; Talmour-Barkan et al., 2022).
Spatial and Single-Cell Technologies Reveal Hidden Biology
Single-cell sequencing has advanced cardiac biology by identifying disease-associated cellular programmes, but tissue dissociation removes spatial context, limiting links between molecular findings and anatomy. Spatial transcriptomics addresses this by measuring gene expression within intact tissue architecture. Integrated spatial and single-cell analyses of myocardial infarction map region-specific inflammation, fibrosis, and remodelling, while cardiomyopathy studies show how localised inflammatory signalling (e.g., endothelial TNFα activation) drives disease progression (Kuppe et al., 2022; Lee et al., 2025).
By bridging molecular programmes directly to anatomical structure, spatial approaches enable anatomically resolved biomarker discovery, improved risk stratification, and more personalised therapeutic strategies in cardiovascular disease. Nevertheless, variability in tissue processing, platform chemistry, analytical pipelines, and cross-study integration continues to challenge reproducibility and consistent clinical translation across cohorts and institutions.
Multi-Omics and Multi-Modal Integration in Clinical Practice
Proteomic and metabolomic profiling capture downstream biological activity that often correlates more directly with clinical outcomes than genetic data alone, enabling earlier intervention and adaptive monitoring, and prompt identification of treatment response or futility.
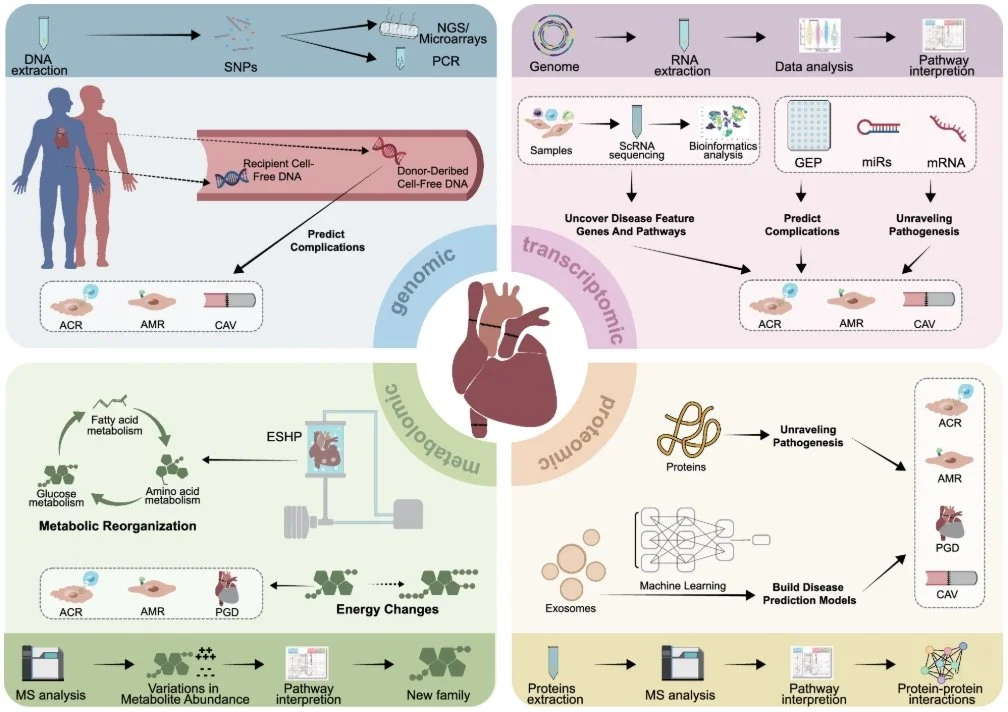
In heart transplantation, multi-omics improves post-transplant monitoring by capturing complementary molecular layers of allograft injury and immune activation that are not fully reflected by histology alone (Zou et al., 2025) (Figure 1). Genomic markers such as donor-derived cell-free DNA provide a quantitative measure of graft injury, while transcriptomic signatures and circulating microRNAs reflect immune and endothelial activation states associated with rejection and disease progression (Singh et al., 2015; Agbor-Enoh et al., 2021).
Figure 1. Multi-omics approaches in heart transplantation.
Schematic overview of genomic, transcriptomic, proteomic, and metabolomic analyses applied to heart transplantation. Genomic profiling of donor- and recipient-derived cell-free DNA and SNPs supports prediction of post-transplant complications. Transcriptomic analyses include single-cell RNA sequencing alongside gene expression, mRNA, and miRNA profiling to identify disease-associated genes and pathways. Proteomic and metabolomic workflows use mass spectrometry–based analyses to investigate pathogenesis, metabolic reprogramming, and energy changes. Integrated multi-omics data contribute to prediction and characterisation of acute cellular rejection (ACR), antibody-mediated rejection (AMR), primary graft dysfunction (PGD), and cardiac allograft vasculopathy (CAV). Adapted from Zou et al., 2025.
Machine learning approaches integrating multi-omic, imaging, and clinical datasets are beginning to reshape cardiovascular research and care pathways. Deep learning models leveraging large-scale transcriptomic and multi-omic data have identified regulators of cardiomyopathy and post–myocardial infarction remodelling with clear translational relevance (Iborra-Egea et al., 2021; Ren et al., 2023).
Integrated omics–ML frameworks have also demonstrated improved predictive performance over conventional clinical risk scores in chronic coronary disease, heart failure, and cardiac transplantation, supporting more informed planning of follow-up, rehabilitation, and advanced interventions (Wallentin et al., 2021; Ouwerkerk et al., 2023; Halloran et al., 2024).
Despite increasing methodological maturity, real-world influence on research and development decisions remains constrained by inconsistent data availability, misalignment between molecular sampling and clinical workflows, and challenges integrating multimodal outputs into regulatory and clinical decision frameworks. Consequently, current implementations remain largely adjunctive, although these approaches collectively demonstrate strong potential to advance predictive, preventive, and personalised medicine (PPPM) in cardiovascular practice (Lin et al., 2025).
Infrastructure, Validation, and Clinical Deployment
The central challenge in precision cardiology is no longer data generation but responsible integration and clinical deployment. As a result, the limiting factor is increasingly the ability to make complex biological data usable at the moment research and early development decisions are made. Without effective integration, translational progress stalls, trials remain heterogeneous, and biologically informed care remains unrealised.
Effective implementation requires harmonised data acquisition, interoperability across imaging and molecular platforms, prospective validation, and regulatory alignment (Țapoș et al., 2026). Decision support frameworks and real-world evidence are essential to translate multi-omic insights into research, development, and translational settings while maintaining safety, transparency, and equity.
Toward Integrated Precision Cardiovascular Care
DNA-based biomarkers remain fundamental for understanding inherited risk; however, DNA-first approaches are insufficient for dynamic cardiovascular disease driven by evolving cellular and spatial biology. Moving beyond single molecular layers toward coordinated multi-omic integration offers a path toward truly mechanism-informed precision cardiovascular medicine.
Supported by robust governance, validation, and interdisciplinary collaboration, this approach improves stratification and informs therapeutic development. As cardiovascular research increasingly confronts biological complexity, integrated multi-omic strategies are shifting from innovation to a necessity in modern precision cardiology.”
AXIA Medicine operates within the constraints of biological complexity, longitudinal data, and decision timing, providing the foundational infrastructure that enables multi-omic and spatial data to inform clinical development beyond anatomical snapshots.
References
Agbor‐Enoh, S., Shah, P., Tunc, I., Hsu, S., Russell, S.D., Feller, E., Shah, K.B., Rodrigo, M.E., Najjar, S.S., Kong, H., Pirooznia, M., Fideli, U., Bikineyeva, A., Marishta, A., Bhatti, K., Yang, Y., Mutebi, C., Yu, K., Jang, M. K., and Marboe, C.C. (2021). Cell-Free DNA to Detect Heart Allograft Acute Rejection. Circulation, 143(12), pp.1184–1197. doi:https://doi.org/10.1161/circulationaha.120.049098.
Halloran, P.F., Reeve, J., Mackova, M., Madill-Thomsen, K.S., Demko, Z., Olymbios, M., Campbell, P., Melenovsky, V., Gong, T., Hall, S. and Stehlik, J. (2024). Comparing Plasma Donor–derived Cell-free DNA to Gene Expression in Endomyocardial Biopsies in the Trifecta-Heart Study. Transplantation, 108(9), pp.1931–1942. doi:https://doi.org/10.1097/tp.0000000000004986.
Henry, A., María Gordillo-Marañón, Finan, C., Schmidt, A.F., João Pedro Ferreira, Karra, R., Johan Sundström, Lind, L., Johan Ärnlöv, Faiez Zannad, Anders Mälarstig, Hingorani, A.D. and R. Thomas Lumbers (2022). Therapeutic Targets for Heart Failure Identified Using Proteomics and Mendelian Randomization. Circulation, 145(16), pp.1205–1217. doi:https://doi.org/10.1161/circulationaha.121.056663.
Iborra-Egea, O., Gálvez-Montón, C., Prat-Vidal, C., Roura, S., Soler-Botija, C., Revuelta-López, E., Ferrer-Curriu, G., Segú-Vergés, C., Mellado-Bergillos, A., Gomez-Puchades, p., Gastelurrutia, P., and Bayes-Genis, A., (2021). Deep Learning Analyses to Delineate the Molecular Remodeling Process after Myocardial Infarction. Cells, 10(12), pp.3268–3268. doi:https://doi.org/10.3390/cells10123268.
Jeong, H.-E., Kim, K.-H., Kim, Y., Yu, H.-Y., Shin, D.M., Kim, O.-H., Kim, B.-J., Park, M.-H. and Kim, J. (2025). Unraveling the cellular characteristics of cardiomyopathy with rare variant-driven gene signatures using multi-omics analysis. Scientific Reports. [online] doi:https://doi.org/10.1038/s41598-025-33901-y.
Kuppe, C., Ramirez Flores, R.O., Li, Z., Hayat, S., Levinson, R.T., Liao, X., Hannani, M.T., Tanevski, J., Wünnemann, F., Nagai, J.S., Halder, M., Schumacher, D., Menzel, S., Schäfer, G., Hoeft, K., Cheng, M., Ziegler, S., Zhang, X., Peisker, F. and Kaesler, N. (2022). Spatial multi-omic map of human myocardial infarction. Nature, 608(7924), pp.766–777. doi:https://doi.org/10.1038/s41586-022-05060-x.
Lee, S.E., Joo, J.H., Hwang, H.S., Chen, S., Evans, D., Lee, K.Y., Kim, K., Hyun, J., Kim, M., Lee, J.S. and Torkamani, A. (2025). Spatial transcriptional landscape of human heart failure. European Heart Journal, 46(31), pp.3098–3114. doi:https://doi.org/10.1093/eurheartj/ehaf272.
Lin, M., Guo, J., Gu, Z., Tang, W., Tao, H., You, S., Jia, D., Sun, Y. and Jia, P. (2025). Machine learning and multi-omics integration: advancing cardiovascular translational research and clinical practice. Journal of Translational Medicine, [online] 23(1). doi:https://doi.org/10.1186/s12967-025-06425-2.
Liu, C.-F., Ni, Y., Moravec, C.S., Morley, M., Ashley, E.A., Cappola, T.P., Margulies, K.B. and Tang, W.H.W. (2021). Whole-Transcriptome Profiling of Human Heart Tissues Reveals the Potential Novel Players and Regulatory Networks in Different Cardiomyopathy Subtypes of Heart Failure. Circulation: Genomic and Precision Medicine, 14(1). doi:https://doi.org/10.1161/circgen.120.003142.
Mora Massad, K., Dai, Z., Petrache, I., Ventetuolo, C.E. and Lahm, T. (2025). Lung endothelial cell heterogeneity in health and pulmonary vascular disease. American Journal of Physiology-Lung Cellular and Molecular Physiology, 328(6), pp.L877–L884. doi:https://doi.org/10.1152/ajplung.00296.2024.
Örd, T., Õunap, K., Stolze, L.K., Aherrahrou, R., Nurminen, V., Toropainen, A., Selvarajan, I., Lönnberg, T., Aavik, E., Ylä-Herttuala, S., Civelek, M., Romanoski, C.E. and Kaikkonen, M.U. (2021). Single-Cell Epigenomics and Functional Fine-Mapping of Atherosclerosis GWAS Loci. Circulation Research, 129(2), pp.240–258. doi:https://doi.org/10.1161/circresaha.121.318971.
Ouwerkerk, W., Pereira, J.P.B., Maasland, T., Emmens, J.E., Figarska, S.M., Tromp, J., Koekemoer, A.L., Nelson, C.P., Nath, M., Romaine, S. P. R., Cleland, J. G. F., Zannad, F., van Veldhuisen, D.J., Lang, C.C., Ponikowski, P., Filippatos, G., Anker, S., Metra, M., Dickstein, K. and Ng, L.L. (2023). Multiomics Analysis Provides Novel Pathways Related to Progression of Heart Failure. Journal of the American College of Cardiology, 82(20), pp.1921–1931. doi:https://doi.org/10.1016/j.jacc.2023.08.053.
Ramón y Cajal, S., Sesé, M., Capdevila, C., Aasen, T., De Mattos-Arruda, L., Diaz-Cano, S.J., Hernández-Losa, J. and Castellví, J. (2020). Clinical implications of intratumor heterogeneity: challenges and opportunities. Journal of Molecular Medicine, 98(2), pp.161–177. doi:https://doi.org/10.1007/s00109-020-01874-2.
Ren, Z.-H., You, Z., Zou, Q., Yu, C.-Q., Ma, Y., Guan, Y.-J., You, H.-R., Wang, X. and Pan, J. (2023). DeepMPF: deep learning framework for predicting drug–target interactions based on multi-modal representation with meta-path semantic analysis. Journal of Translational Medicine, 21(1). doi:https://doi.org/10.1186/s12967-023-03876-3.
Singh, N., Heggermont, W., Fieuws, S., Vanhaecke, J., Van Cleemput, J. and De Geest, B. (2015). Endothelium-enriched microRNAs as diagnostic biomarkers for cardiac allograft vasculopathy. The Journal of Heart and Lung Transplantation, 34(11), pp.1376–1384. doi:https://doi.org/10.1016/j.healun.2015.06.008.
Talmor-Barkan, Y., Bar, N., Shaul, A.A., Shahaf, N., Godneva, A., Bussi, Y., Lotan-Pompan, M., Weinberger, A., Shechter, A., Chezar-Azerrad, C., Arow, Z., Hammer, Y., Chechi, K., Forslund, S.K., Fromentin, S., Dumas, M.-E., Ehrlich, S.D., Pedersen, O., Kornowski, R. and Segal, E. (2022). Metabolomic and microbiome profiling reveals personalized risk factors for coronary artery disease. Nature Medicine, 28(2), pp.295–302. doi:https://doi.org/10.1038/s41591-022-01686-6.
Țapoș, G.-F., Cîmpeanu, I.-A., Predescu, I.-A., Liga, S., Păcurar, A.T., Vlad, D., Boru, C., Luca, S., Crișan, S., Văcărescu, C. and Luca, C.T. (2026). An Integrative Review of the Cardiovascular Disease Spectrum: Integrating Multi-Omics and Artificial Intelligence for Precision Cardiology. Diseases, [online] 14(1), p.31. doi:https://doi.org/10.3390/diseases14010031.
Wallentin, L., Eriksson, N., Olszowka, M., Grammer, T.B., Hagström, E., Held, C., Kleber, M.E., Koenig, W., März, W., Stewart, R.A.H., White, H.D., Åberg, M. and Siegbahn, A. (2021). Plasma proteins associated with cardiovascular death in patients with chronic coronary heart disease: A retrospective study. PLoS Medicine, [online] 18(1), p.e1003513. doi:https://doi.org/10.1371/journal.pmed.1003513.
Wang, R.-S., Maron, B.A. and Loscalzo, J. (2023). Multiomics Network Medicine Approaches to Precision Medicine and Therapeutics in Cardiovascular Diseases. Arteriosclerosis, Thrombosis, and Vascular Biology, 43(4), pp.493–503. doi:https://doi.org/10.1161/atvbaha.122.318731.
World Health Organization (2025). Cardiovascular diseases (CVDs). [online] World Health Organization . Available at: https://www.who.int/news-room/fact-sheets/detail/cardiovascular-diseases-(cvds).
Zou, Z., Han, J., Zhu, Z., Zheng, S., Xu, X. and Liu, S. (2025). Transforming heart transplantation care with multi-omics insights. Journal of Translational Medicine, [online] 23(1). doi:https://doi.org/10.1186/s12967-025-06772-0